New Approach to Teaching User Experience Design Skills Wins Award
A team of researchers at the Kahlert School of Computing has won a best paper honorable mention award for their paper describing a new approach to teaching students about design empathy in the classroom. The award-winning paper will be presented at the Designing Interactive Systems (DIS) 2024 conference taking place this week in Copenhagen, Denmark.
When software companies like Apple, Google, and Microsoft create or improve applications they start by doing lots of work to better understand their users. User experience researchers collect data: they observe, interview, survey, and study potential users so that they can understand how the software might fit into their lives, also known as developing “design empathy” for those potential users. Having design empathy is valuable even to other people on the software teams who are not designers – software engineers and project managers make better software when they understand more about who is using it and how it will be used.
This is even more crucial when people are creating software for people whose lives are very different from their own: for example, when someone who does not have a disability is designing software that will be used by people who do have disabilities, as is often the case. However, teaching students about design empathy is a difficult challenge for professors: collecting this data is itself a skill, and it requires the participation of other third parties to serve as “the users.” Again, when working with a population of users with disabilities, this can be a particular challenge.
Newly graduated doctoral students Tamanna Motahar and Noelle Brown, along with their advisors Professors Eliane Stampfer Wiese and Jason Wiese, developed a new approach to more effectively teach design empathy in the classroom. The approach leverages the fact that many people post publicly about their experiences on social media platforms such as Reddit. The team first collected some of these posts for a particular user community: people who have had a spinal cord injury. They categorized each post based on its subject, and then used those categories to guide the creation of fictional design scenarios: going to the grocery store, going on vacation, returning to school after an injury, and handling air travel.




They then selected some of the posts, paraphrased or changed their wording to protect the identity and privacy of the original poster, and curated the posts into a reading assignment that would accompany each scenario. Finally, they wrote a multi-part assignment around these scenarios and posts. In part 1, students are each assigned to think through one of the design scenarios using a series of questions to prompt their thinking. Part 2 involves reading the curated and paraphrased posts, described above. In part 3 the students come together in class with groups of 3-4 students who were assigned different scenarios to discuss their thoughts after reading the posts. In part 4, the students revisit their individual design scenarios from part 1 to see if they would add or change anything.
To test the idea, the team deployed the project in a small class that Professor Jason Wiese was teaching. “What we saw was that despite the students providing thoughtful responses to the design prompt in part 1, their in-class conversations and final responses in part 4 showed that their perspective really shifted and they considered the impact of many more real-world factors after reading the posts,” said Motahar, the lead author of the paper.
The research team plans to further refine the assignment and to work towards sharing the assignment materials more broadly, so that other teachers can use them to teach students about developing design empathy. One important consideration in this work, and especially about sharing the assignment materials, is about the ethical implications of using social media posts for a purpose other than what was originally intended by the poster, and also to protect the privacy of those posters. The research team followed broadly agreed upon best practices for working with such social media data, including removing any identifiers - including usernames or locations - and changing the words of the posts before using them while still preserving the meaning of the original post.
The full paper is available online: Tamanna Motahar, Noelle Brown, Eliane Stampfer Wiese, and Jason Wiese. Toward Building Design Empathy for People with Disabilities Using Social Media Data: A New Approach for Novice Designers. In Designing Interactive Systems Conference (DIS '24). https://doi.org/10.1145/3643834.3660687
Prof. Patil Selected as Faculty Co-Director of the University of Utah’s DATASET Initiative
Associate Prof. Sameer Patil from the Kahlert School of Computing has been selected as a faculty co-director of the Data Science & Ethics of Technology (DATASET) Initiative at the University of Utah to serve alongside co-director Prof. Manish Parashar. Prof. Patil replaces past co-director Prof. Olivia Sheng who is leaving the University of Utah this summer to join the W. P. Carey School of Business at Arizona State University.

The DATASET Initiative is a part of the One Utah Data Science Hub, a university-wide effort designed to enhance research and infrastructure in data science and data-enabled science. DATASET was launched in the fall of 2022 to engage in foundational questions about the role of data in science and society. The initiative aims to bring together research and expertise in the theoretical, technical, ethical, historical, and policy/legal dimensions of data across the University of Utah. The goals of the DATASET initiative are well-aligned with Prof. Patil's courses on Ethics in Data Science and Human Aspects of Security and Privacy. “I am honored to have been selected for the role of co-director and excited to leverage the position for promoting data sharing and open science to facilitate transparency, verifiability, and replication of data-driven research results,” Patil said.
In his role as co-director, Prof. Patil will help develop the vision and specific activities of the DATASET initiative and engage with the DATASET Advisory Board. Associate Director of the One Utah Data Science Hub Penny Atkins said, "We were impressed by Prof. Patil's experiences, perspectives, and goals related to DATASET and believe that he will be an asset to our initiative going forward."
Kahlert School Research on Visualizations for Metabolic Networks Published in Nature Cell Biology and Science
In September 2019, Jordan Berg, then a PhD student in the lab of Professor Jared Rutter in the Biochemistry Department, contacted Professor Bei Wang Phillips in the Kahlert School of Computing and the Scientific Computing and Imaging Institute regarding how best to design a tool for the analysis and visualization of metabolic networks. Over the next four years, then PhD student Youjia Zhou from Professor Wang Phillips’ lab collaborated with Jordan and Professor Rutter’s team. The collaboration has contributed toward two recent prestigious publications: one in Nature Cell Biology led by Jordan (Jordan A. Berg et al., 2023) and another in Science led by Dr. Kevin G. Hicks (Kevin G. Hicks et al., 2023). Berg and Zhou defended their PhDs in the Springs of 2022 and 2023, respectively. Dr. Zhou’s PhD thesis is titled “Topology-Based Visualization of Graphs and Hypergraphs.”
Metabolism is an important process in our bodies that affects various aspects of our health, such as cell growth and development, stress response, and energy production. It is a complex, interdependent network, and small changes in one area can have widespread effects throughout the entire metabolic system. Existing tools for analyzing metabolic networks suffer from various problems: they may be outdated, have limited functionality, be too technical to run for a biochemist, or completely unusable. To address these shortcomings, Zhou and Wang Phillips collaborated with Berg and Rutter’s team to develop new tools that leverage interactive graph visualizations. These tools are designed to analyze complex metabolic networks with a focus on detecting patterns and studying interactions between proteins and metabolites.


“Our vision is to develop practical visualizations of networks that help domain scientists with their knowledge discovery process,” said Professor Wang Phillips.
Pattern Recognition
The first interactive tool is called Metaboverse, a user-friendly tool that helps with the exploration of metabolic networks. In particular, it enables pattern recognition from a large, varied library of possible metabolic patterns. In the context of metabolism, a reaction pattern refers to a metabolic reaction that shows some change between the inputs and outputs of the reaction. Metaboverse enhances the understanding of detected patterns by providing a dynamic exploratory interface, which helps domain scientists come up with new ideas.
Previously, scientists searched for desired patterns manually, which was time-consuming and incomplete. Similarly, other available tools could find and identify only a few types of patterns and had the potential to overlook crucial insights. Metaboverse helps solve these problems: it introduces a broad collection of patterns and, using automated techniques, provides a quick and accurate way for scientists to detect complex patterns that they might have previously missed.
In terms of practical applications, Metaboverse was used to find patterns in the metabolism of early-stage human lung adenocarcinomas (LUAD), a type of lung cancer. The goal was to identify which metabolites could be used to diagnose the disease at an early stage. Metaboverse was able to prioritize patterns in nucleotide metabolism reliably; consistent with the original study as well as the manual re-analysis of the data using other existing tools. In addition, Metaboverse helped discover new patterns related to xanthine metabolism and a group of reactions linked to a decrease in lysine. These discoveries were made using data from a previous study where changes in these metabolites were observed but not fully explored.
Analysis of Protein-Metabolite Interactions
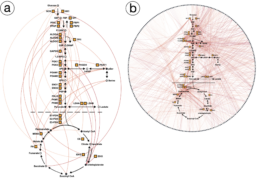
Protein-metabolite Interactions (PMIs) indicate how proteins and metabolites communicate metabolic status to different cellular processes. In metabolic networks, finding PMIs that mediate these networks can be difficult because they often have low affinity. To help solve this problem, Zhou and Wang Phillips worked with Berg and Rutter’s team to create MIDAS, a library that helps find PMIs systematically. Specifically, the researchers helped develop Electrum, a flexible, user-friendly visualization portal that aids in the understanding and analysis of the PMI data generated by MIDAS. While other general visualization platforms are unable to handle PMI data because of the specific analytical capabilities it requires, Electrum can differentiate between known and unknown PMIs, which can lead to new discoveries.
The usefulness of Electrum was demonstrated through a case study in which interactions within and between pathways in carbohydrate metabolism were discovered. Carbohydrate metabolism enzymes drive most of the energy production and biosynthetic precursor generation of cells, and their regulation involves metabolite interactions. The MIDAS platform identified 830 putative PMIs, many of which were previously unknown. Electrum provided interactive visualizations to help domain scientists comprehend the PMIs within and between pathways, and the visual analytics results revealed regulation within and between pathways in carbohydrate metabolism. The collaboration led to a recent publication in Science led by Dr. Kevin G. Hicks.
In the future, the researchers will continue developing and enhancing interactive visualization tools to help domain scientists better explore and understand complex networks.
Award-winning Research from the Kahlert School Examines Student Understanding of Code Quality
Kahlert School of Computing Ph.D. student Sara Nurollahian and Assistant Prof. Eliane Wiese have received the best paper award for the Software Engineering Education and Training Track at the International Conference on Software Engineering (ICSE 2023) for their paper titled Improving Assessment of Programming Pattern Knowledge through Code Editing and Revision that was presented at the conference earlier in May 2023.
When programmers write code, it’s not enough for a program to run, or even to work, as expected. Even if a computer is capable of parsing and properly executing the code, humans need to be able to read and understand it as well in order to maintain, test, and debug it over the course of its life. Given the complex nature of modern code, how can computer science students learn to think about, design, and write quality code that is highly technical and highly readable? Addressing this larger question is the motivation behind the research in the award-winning paper. Advised by Prof. Eliane Wiese, Nurollahian examined how student coders demonstrate understanding of their code through different activities. “ICSE is the premier conference on software engineering and it’s exciting to have Sara present her work in their education track, which reaches a broad range of Computer Science education researchers around the world,” Prof. Eliane Wiese said.


In their research, Nurollahian and Wiese used a survey to measure student understanding of two important code quality issues: avoiding repetitions and returning boolean expressions rather than literals in simple methods. These types of quality issues are similar to simplifying math expressions. For example, it is difficult to think of the two integer values for x and y for the equation 4x + 8y + 3 = 39, but it is a lot easier for this equivalent one: x + 2y = 9.
A common approach to research in this area is to measure student' understanding by examining the code they write. However, students may write elegant code without understanding the principles behind it or write poor-quality code even if they know better. Instead of looking only at code writing, the study additionally asked students to revise their own code and edit some code written by others. For code writing, students were given the signature of a method along with a description of its desired behavior and asked to fill in the body. For editing, students were given low-quality code blocks written by others and asked to improve them. Similarly, for revising, students who wrote low-quality code in the writing task were shown their own code and asked to improve it. However, for revising, students were given a series of hints, starting with a simple prompt to revise (without indicating what was wrong or how to fix it), and ending with an example for a similar code block.
“These three tasks allowed us to examine the revising and editing performance of students who wrote low-quality code, considering that students whose code is low in quality due to knowledge gaps will not be able to perform well in revising and editing without receiving extra support, ” Nurollahian said. Students who wrote poor-quality code needed different levels of hints to revise successfully, allowing the researchers to differentiate between knowledge gaps and other factors, such as motivation. For example, over half of the students who returned boolean literals in the writing task were able to revise successfully when prompted to improve (without getting information on what to fix or how), suggesting that they simply needed more motivation. However, the majority of the students who wrote code that contained repetitions were not able to revise successfully, even after seeing hints on what to change and an example of implementing the change in a similar code block, indicating that they needed greater knowledge in this area. The editing tasks additionally showed that code writing can give an incomplete picture of student understanding. While students who wrote well initially were generally able to edit more successfully, there were exceptions in both directions. For returning boolean literals (vs. expressions), between one quarter and one third of students who wrote poorly were successful in editing someone else's code, while a similar percentage of students who initially wrote well made unsuccessful edits. For code repetitions, fewer students overall wrote or edited successfully, but the general pattern still held.
These findings highlight the benefit of using a variety of tasks to measure student understanding of code quality. The work also underscores the importance of further research to examine the quality issues that are challenging for students. As Nurollahian explained, “The distinction between simple and complex quality issues can help instructors allocate their time and effort to the more challenging ones. Additionally, distinguishing between students who have deep conceptual gaps and those whose low-quality code is due to other reasons, such as lack of motivation, can help appropriately tailor instruction, feedback, and learning interventions.”
Kahlert School of Computing Faculty and Students Heading to Hamburg for CHI 2023
The Kahlert School of Computing is heading to Hamburg, Germany for the CHI 2023 conference, which will be held from April 23rd - April 28th. Considered to be the leading global conference on Human-Computer Interaction, the ACM CHI Conference on Human Factors in Computing Systems brings together business and academic leaders to discuss ways to develop and enhance new interactive digital technologies.
This year, the Kahlert School of Computing will have particularly strong representation at CHI with seven papers and one case study. The paper Never Skip Leg Day Again: Training the Lower Body with Vertical Jumps in a Virtual Reality Exergame by Kahlert School of Computing faculty affiliate Jens Harald Kreuger is being honored with the Best Paper Honorable Mention, a selective honor awarded to only the top 5% of papers from nearly 3,200 submissions. Overall, the University of Utah will be represented by 14 individuals including six professors alongside a mix of researchers and students. "The strong presence of the Kahlert School of Computing at the world’s leading HCI conference is a testament to the school’s growing emphasis on a human-centered approach to designing technologies of the future," said Prof. Mary Hall, Director of the Kahlert School of Computing.
Faculty and students from the Kahlert School of Computing will be presenting seven papers on a variety of subjects ranging from the impact of smart home technology on rehabilitation centers to methods used to spread misinformation through data visualization:
Jason Wiese (University of Utah), John R. Lund (University of Utah), and Kazi Sinthia Kabir (University of Utah)
Data Abstraction Elephants: The Initial Diversity of Data Representations and Mental Models
Kay P. Williams (University of Arizona), Alex Bigelow (Stardog), and Katherine E. Isaacs (University of Utah).
Joshua Dawson (University of Utah), Thomas Kauffman (University of Utah), and Jason Wiese (University of Utah)
Misleading Beyond Visual Tricks: How People Actually Lie with Charts
Maxim Lisnic (University of Utah), Cole Polychronis (University of Utah), Alexander Lex (University of Utah), and Marina Kogan (University of Utah)
Never Skip Leg Day Again: Training the Lower Body with Vertical Jumps in a Virtual Reality Exergame
Sebastian Cmentowski (University of Duisburg-Essen), Sukran Karaosmanoglu (Hamburg University), Lannart E. Nacke (University of Waterloo), Frank Steinicke (Hamburg University), and Jens Krüger (University of Duisburg-Essen, University of Utah)
🏆 BEST PAPER HONORABLE MENTION
Emma Ning (University of Illinois at Chicago), Andrea T. Cladek (University of Illinois at Chicago), Mindy K. Ross (University of Illinois at Chicago), Sarah Kabir (University of Illinois at Chicago), Amruta Barve (University of Illinois at Chicago), Ellyn Kennelly (Wayne State University), Faraz Hussain (University of Illinois at Chicago), Jennifer Duffecy (University of Illinois at Chicago), Scott Langenecker (University of Utah), Therea Nguyen (University of Illinois at Chicago), Theja Tulabandhula (University of Illinois at Chicago), Olusola A. Ajilore (University of Illinois at Chicago), Alexander P. Demos (University of Illinois at Chicago), and Alex Leow (University of Illinois at Chicago)
Troubling Collaboration: Matters of Care for Visualization Design Study
Derya Akbaba (Linköping University), Devin Lange (University of Utah), Michael Correl (Tableau Software), Alexander Lex (University of Utah), and Miriah Meyer (Linköping University)
"The strong presence of the Kahlert School of Computing at the world’s leading HCI conference is a testament to the school’s growing emphasis on a human-centered approach to designing technologies of the future."
Prof. Mary Hall, Director of the Kahlert School of Computing.
In addition to the seven papers, Postdoc researcher Johanna Cohoon will present a case study titled Adapting to Challenges in Qualitative Fieldwork through Theoretical Sampling co-authored with James Howison of the University of Texas at Austin.
In addition, students and faculty from the school will participate and present their work in the Symposium on HCI Education (EduCHI) and the Workgroup on Interactive Systems in Healthcare (WISH) workshop held at the conference.
EduCHI is the premier venue focused on research and practice connected to teaching HCI and supporting community building among HCI instructors.
- Building "Design Empathy" for People with Disabilities: An Unsolved Challenge in HCI Education
Tamanna Motahar (University of Utah), Noelle Brown (University of Utah), Eliane Wiese (University of Utah), and Jason Wiese (University of Utah) - Structured Self-Reflection to Evaluate Contextual Inquiry Techniques
Noelle Brown (University of Utah), Nidhi Patel (University of Utah), Xavier Davis (University of Utah), and Eliane Wiese (University of Utah)
WISH aims to foster a community around innovations in consumer and medical health and wellbeing by connecting academic and industry researchers in HCI, medical informatics, health informatics, digital health, and beyond.
- HCI Research Challenges in Complex Healthcare Context (Work in Progress)
Tamanna Motahar (University of Utah), Kazi Sinthia Kabir (University of Utah), Joshua Dawson (University of Utah), Jason Wiese (University of Utah) - The Impact of Spinal Cord Injury on Participation in Human-Centered Research (Research Highlight)
Kazi Sinthia Kabir (University of Utah), Ahmad Alsaleem (University of Utah), and Jason Wiese (University of Utah)