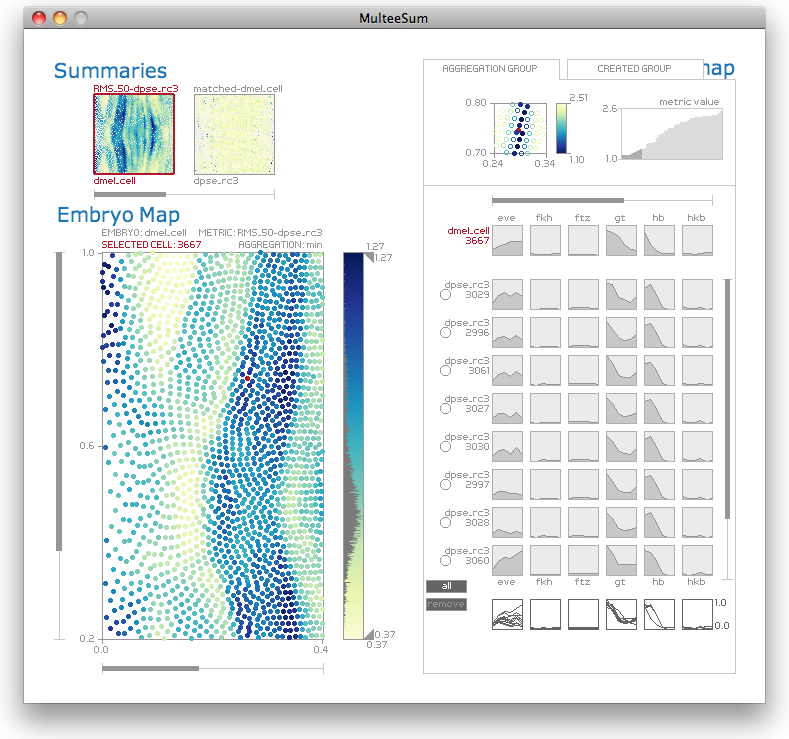
MulteeSum is a visualization system that supports inspection and curation of data sets showing gene expression over time, in conjunction with the spatial location of the cells where the genes are expressed -- it is the first tool to support comparisons across multiple such datasets. MulteeSum is part of a general and flexible framework that allows biologists to explore the results of computations that mix spatial information, gene expression measurements over time, and data from multiple species or organisms. MulteeSum was developed to support the research activities of biologists studying developing Drosophila embryos.
There is a companion paper for this work that describes the design process and decisions that went into the making of MulteeSum. This paper appeared as part of the proceedings of the 2010 IEEE Information Visualization Conference. [paper]